Research areas
Areas of interest and activity.
Go to:Ethnobiology | Bioacoustics |
Ethnobiology
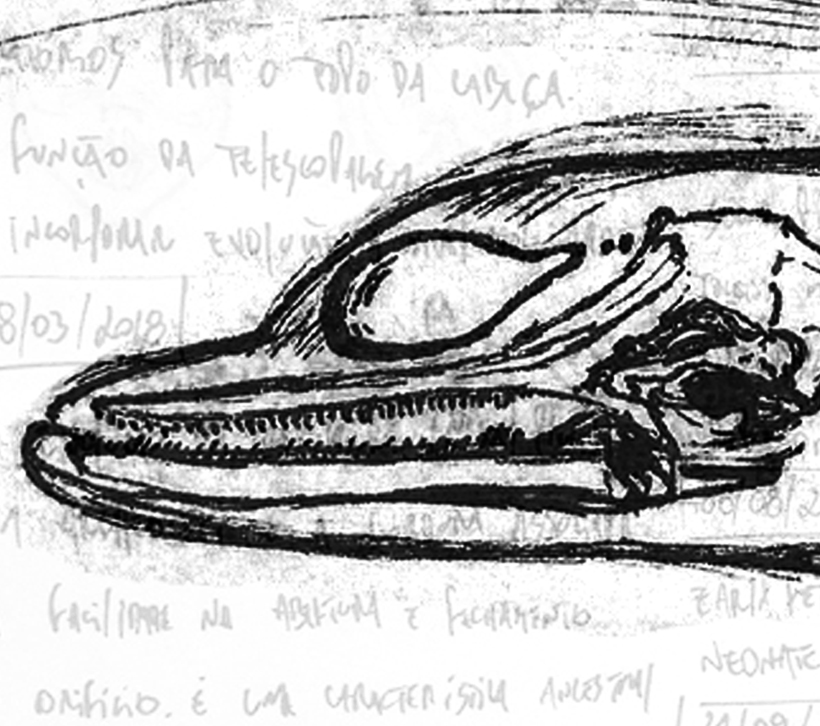
This technique is the oldest scientific discipline of medicine as the first human Histologydissection is documented in Alexandria in the third century B.C. During my masters and PhD, I have basically used gross dissections to investigate the sound-generating structures of some dolphin species. The first species I have studied was the Franciscana dolphin, which I described the development of the main structures involved in sound production, and the general differences between adults and calves. A dissection is basically removing tissues ‘layer by layer, keeping adjacent structures intact as much as possible’ as stated by Schenkkan (1972) while investigating the broadest study comparing the nasal cavity of several dolphin species. The descriptions are based on notes and photographs made during the dissections.
Although I have focused my studies on small dolphins (that for me are the most easy to be studied this way), I have also the opportunity to dissect, literally, from small freshwater fishes to sperm whales. For small fishes, I clear and stained (c&s) some specimens to analyze the skeleton morphology based on Taylor and Van Dyke (1985). With this technique, we remove soft tissues while coloring bones and cartilages with blue and pink, respectively. For big whales, we need a bit more infrastructure. The first time I dissected a whale with all the tools available was in Büsum, Germany, where we had a truck to hold the big pieces of blubber while we were accessing its internal organs. No, we did not could see the lens of the sperm whale, maybe it is just visible using imaging analysis such as CT scans.
Literature Cited
EJ Schenkkan (1972) On the nasal tract complex of Pontoporia blainvillei Gervais and d'Orbigny 1844 (Cetacea, Platanistidae). Investigations on Cetacea
Histology and Anatomic Geometry
I have used histological techniques to assess the morphology of the nasal tract in fetal specimens of some dolphin species using hematoxylin-eosin staining. I have prepared the head of a Franciscana dolphin fetus, and compared to other species already prepared and stored in scientific collections. To demonstrate the morphology of the structures, I have digitized a series of 50 of these slices (i.e., spaced 40 µm each other) and used this image sequence as a input for 3D reconstructions using Slicer3D. In that case I have even converted the image sequence to DICOM format, but you can also use jpeg images as well. The results can be seen in this paper.
I grouped these two approaches here just because I have somehow combined them in terms of my research questions, which was to demonstrate the development of soft-tissue structures throughout the development. The difference is how the sequence of images are generated and how to process them. For late fetuses and very young individuals it is possible to use conventional Magnetic Resonance Imaging (MRI) scans which takes a long time. However, computed tomography (CT) scans demonstrated to be useful to define the fat structures and other connective tissues. The CT-scan is quicker than the MRI, which makes the logistics more productive and less stressful because you always depend on the patience and disponibility of an operator that saves time to scan some dolphins' heads. Since the best scans are placed in human clinics, I put a lot of effort on trying to get access to those ones. The first time I scanned a dolphin I passed through the waiting room carrying a dead dolphin the size of a nine years old kid.
The images can be analyzed, segmented and build 3D of target structures using Slicer3D. Additionally, you can also measure your models with high precision, and prepare it for printing. Not only the 3D models can be stored in digital repositories, but printing these models could be useful for science outreach.
Bioacoustics
I have started working with acoustics properly when I did a training course with Seiche to work as Passive Acoustic Monitoring (PAM) operator. But it was definitely in the postdoc I started digging deep into the topic. I have learned how to set and deploy SoundTraps, Audiomoths, HTI’s attached to a TASCAM recorder and CPOD’s.
Besides the basic modules used for seismic surveys, I have been using PAMGuard for many things such as for localization with four array hydrophones in the boat, whistle detection and classification and Long-Term Spectral Average. I have prepared an algorithm in Python to automate the PAMGuard outputs such as the ‘whistle and moan detector’ and ‘whistle classifier’ modules. The idea behind is to apply a post-processing step to eliminate potential false-positives and determine what I called ‘acoustic encounters’. This code is part of a big project investigating the acoustics and movements of the Indian Ocean humpback dolphin(Sousa plumbea) in South African waters. This is the first step for what I think is ideal to start thinking on automatically identify dolphin signature whistles by its definition (see Janik et al. 2013). Another software I use a lot is Raven, which is useful to annotate sounds and it is free (sometimes it crashes in my laptop, but maybe it is because of my laptop). Very useful!
I am now diggind a bit more signal processing in Python, then using the Scipy and Librosa libraries. I have found an algorithm to perform the same LTSA as Beluga does, but I have found an error and am still not confident on sharing that. For now I am mostly interested in building sound libraries for sound classification using machine learning, so will not go to far for now.